Ridzky Yuda
Welcome
Thank you for visiting my GitHub AcademicPages site. I created this platform to share my research Portfolio, technical expertise, and ongoing projects highlighted in my CV, and to foster collaborations across academia and industry. When time permits, I write short Blogpost discussing recent published works in stem cell biology and immunology into plain language.
About Me
Trained as a stem cell and immunobiologist, I have conducted biomedical research across premier institutions in Europe and the United States, gaining broad expertise in both fundamental and translational science. My research aims to define the molecular mechanisms that regulate stem cell fate decisions and to determine how their disruption contributes to aging-associated dysfunction and hematologic disease. I am particularly interested in how inflammatory stress reshapes hematopoietic stem cell (HSC) commitment during aging (Fig.1).
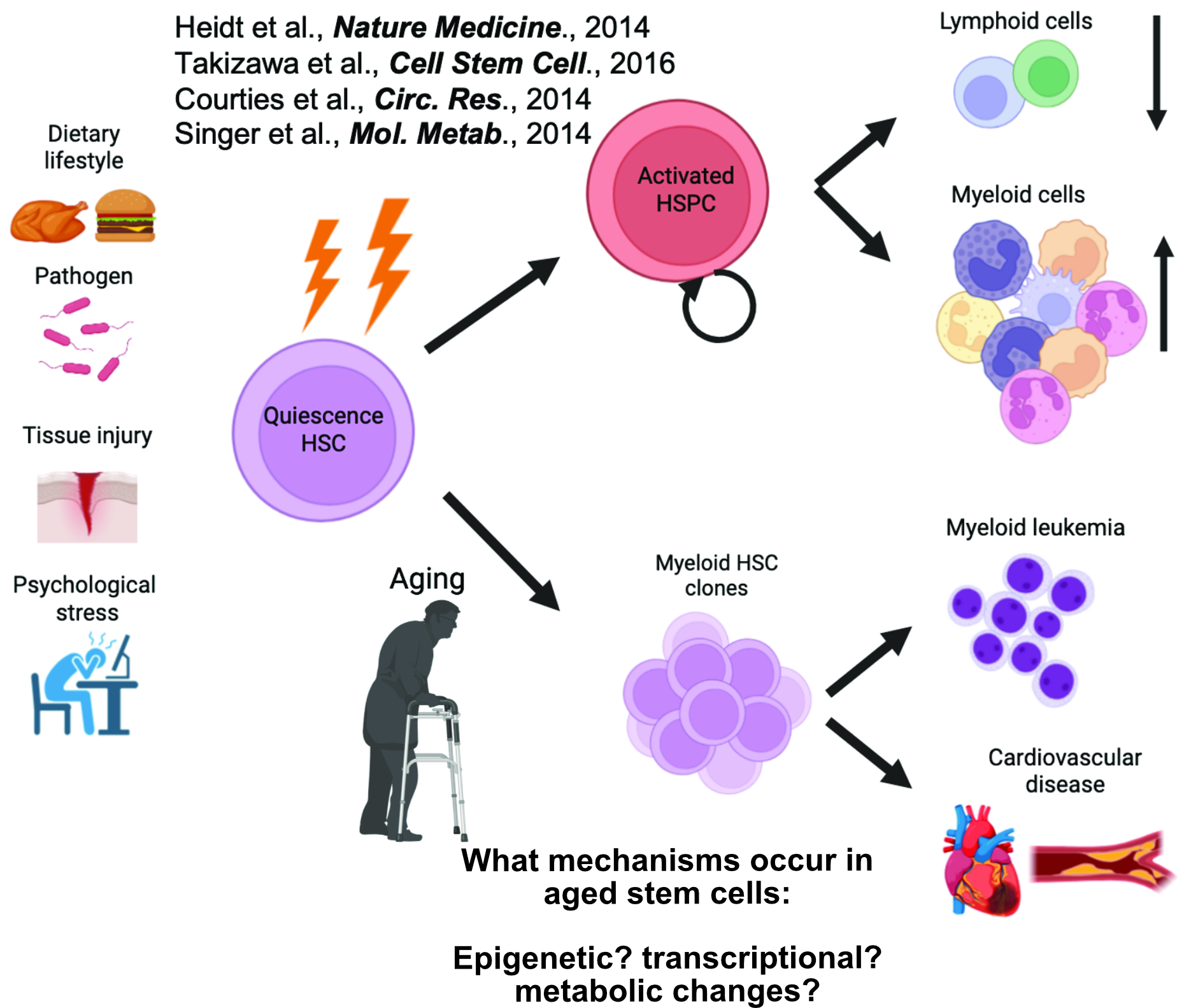 Fig. 1 My long-term research goal is to investigate the impact of age-associated molecular changes in underlying stem cell behavioural changes
Fig. 1 My long-term research goal is to investigate the impact of age-associated molecular changes in underlying stem cell behavioural changes
My scientific endeavour on the biology of stem cell and macrophage aging had started when I did my Ph.D. thesis in the laboratory of prominent HSC and macrophage biologist, Prof. Michael Sieweke, at Technische Universität Dresden, Germany. In Sieweke Lab, I was involved in a collaborative research involving French and German research institutions (Centre d’Immunologie de Marseille-Luminy, Center for Regenerative Therapies Dresden, and the Max Delbrück Center for Molecular Medicine in Berlin) demonstrating that alveolar macrophages—a lung-resident macrophage population—can undergo extensive self-renewal in vitro while maintaining their core cellular identity (Fig. 2A). We further established methods to genetically engineer these cells using lentiviral vectors, expanding their utility for mechanistic and translational studies (Fig. 2B). 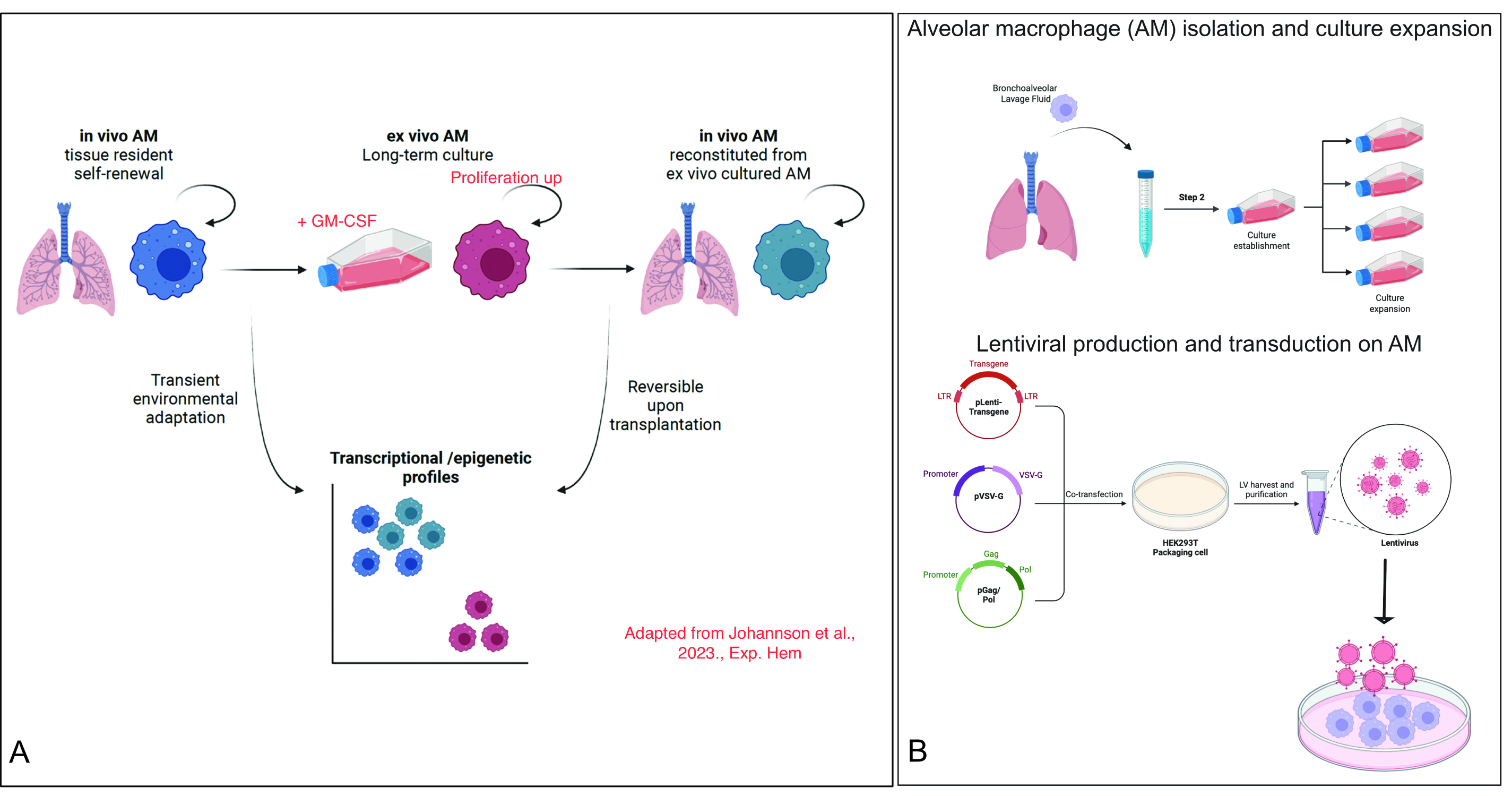 Fig. 2 (A) Epigenetic identity maintenance in self-renewing alveolar macrophages. (B) Methods to genetically manipulate alveolar macrophages.
Fig. 2 (A) Epigenetic identity maintenance in self-renewing alveolar macrophages. (B) Methods to genetically manipulate alveolar macrophages.
In my current postdoctoral project, I integrate in vivo disease modeling with single-cell multi-omics approaches, including scRNA-seq and scATAC-seq, to resolve transcriptional and epigenetic regulatory programs at cellular resolution. I design and execute murine models such as chronic alcohol exposure, LPS-induced inflammation, and bone marrow reconstitution, and I independently develop computational pipelines to analyze and interpret high-dimensional datasets. I found how chronic inflammatory stress alters HSC lineage commitment via sustained inflammation induced by epigenetic derepression of transposable elements (TEs) amplified by aging (Fig. 3).
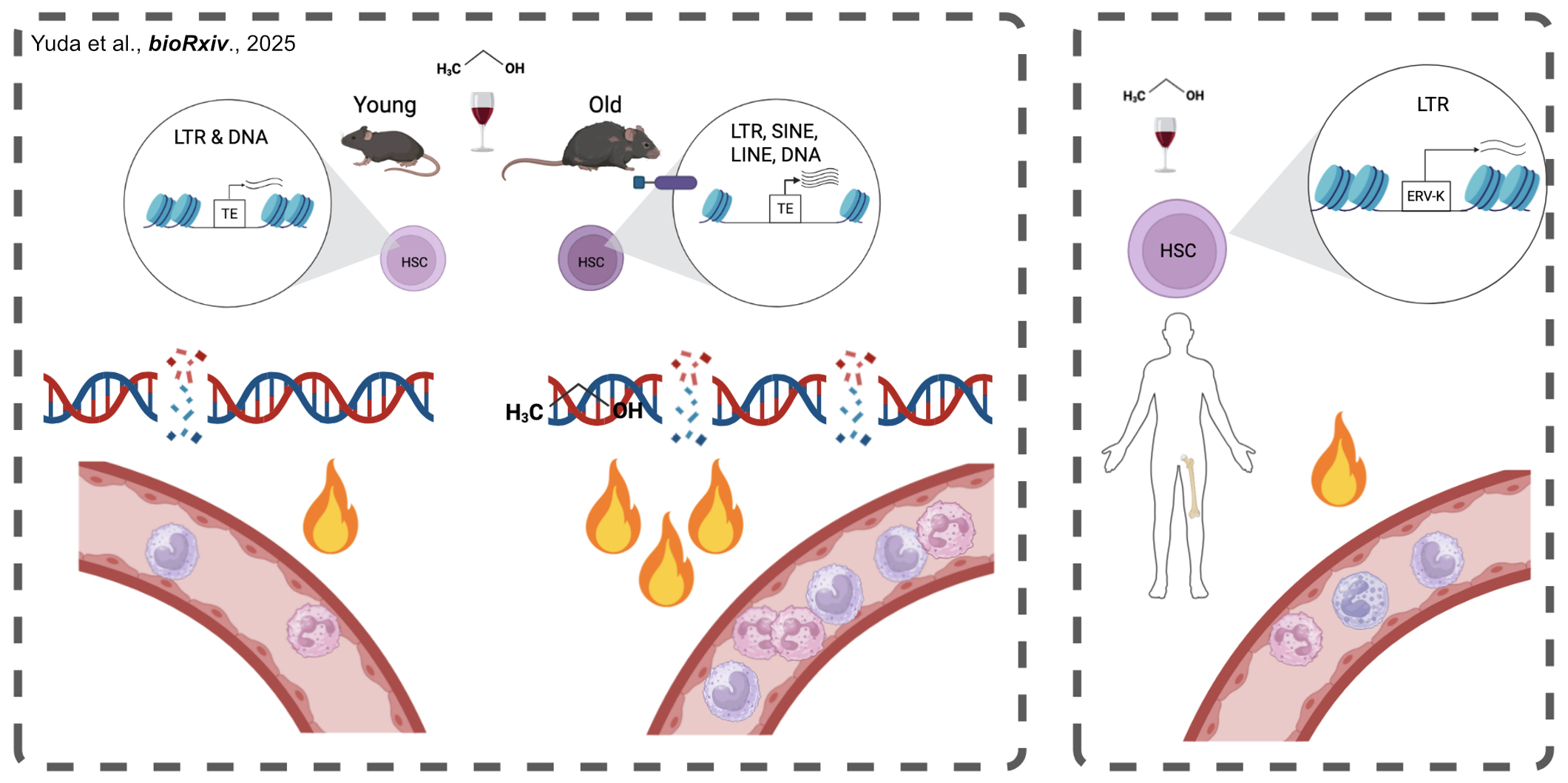 Fig. 3 We found aging amplifies transposable element–associated inflammatory myeloid bias in HSCs under chronic alcohol exposure Yuda et al., bioRXIV., 2025
Fig. 3 We found aging amplifies transposable element–associated inflammatory myeloid bias in HSCs under chronic alcohol exposure Yuda et al., bioRXIV., 2025
For this work, I was awarded an independent and competitive fellowship grant from the Maryland Stem Cell Research Fund (MSCRF) to lead genome-wide epigenomic analysis aimed at defining the mechanisms underlying alcohol-induced inflammation in HSCs. I served as a Principal Investigator (PI), with full responsibility for conceptualization, study design, and execution. Furthermore, my work also extends to neutrophil biology (Fig. 4), where I am currently involved in an ongoing collaboration with the lab of Prof. Lutsenko at Johns Hopkins University by providing my expertise on multiparameter spectral flow cytometry and stem cell isolation.
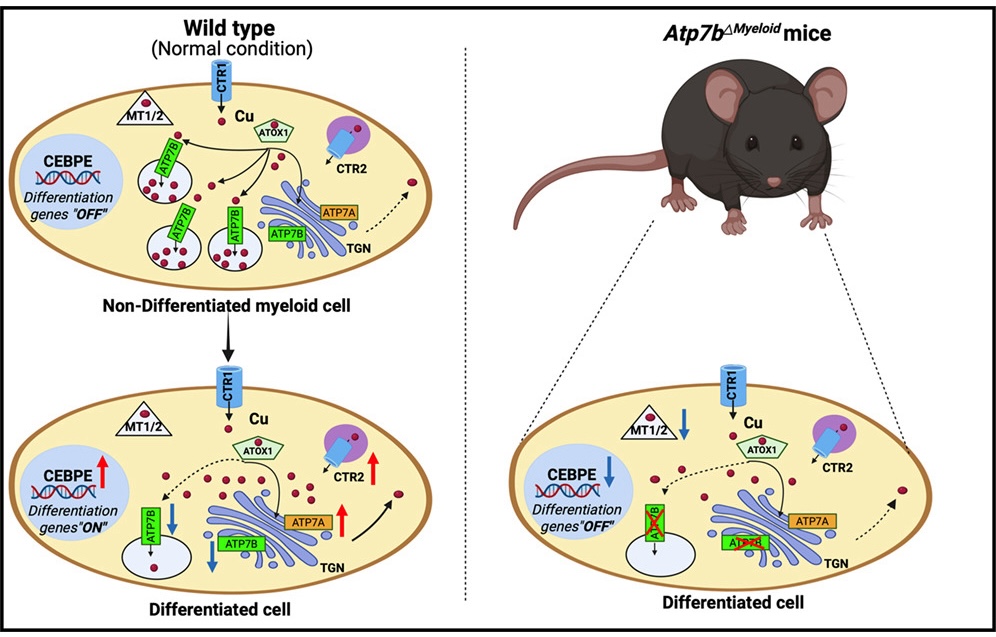 Fig. 4 Neutrophil maturation depends on ATP7B-regulated copper homeostasis Som Dev et al., Cell Reports., 2026
Fig. 4 Neutrophil maturation depends on ATP7B-regulated copper homeostasis Som Dev et al., Cell Reports., 2026
In the meantime, I also developped a newly built R package degTools available in GitHub. This package provides a streamlined wrapper to perform automated differential gene and transposable element expression analyses across cell clusters and data batches/cohorts within a scRNA-seq Seurat object, leveraging Seurat’s FindMarkers function. The package is currently in beta but is available for download and can be readily integrated into analysis workflows.
Overall, my expertise lies at the intersection of experimental hematology, immunology, and integrative genomics. I combine rigorous in vivo experimentation with independent bioinformatic analysis to generate mechanistic insight from complex biological systems.
